15.4 Enzymes
Enzymes, derived from the Greek words En (in) and Zyme (yeast), are biological catalysts. They are complex protein molecules that significantly alter the speed of metabolic activities within living organisms. Enzymes exhibit high specificity in their action and are particularly sensitive to environmental factors like temperature and pH.
15.4.1 Role of Enzymes as Biocatalysts
-
Metabolism: This refers to the intricate series of chemical reactions occurring in living organisms essential for maintaining life. These reactions facilitate growth, reproduction, structural maintenance, and responses to environmental stimuli.
- Anabolic Reactions: Involve the synthesis of larger, more complex molecules from simpler ones (e.g., protein synthesis).
- Catabolic Reactions: Involve the breakdown of larger molecules into smaller ones (e.g., digestion).
- Metabolic reactions fundamentally represent energy transfers, with enzymes playing a critical role in converting chemicals from one state to another. They accelerate biochemical reactions and regulate metabolic pathways.
-
Substrates and Products:
- Substrates: The molecules upon which enzymes act.
- Products: The various molecules formed when enzymes convert substrates.
How Do Enzymes Work?
Enzymes function by temporarily binding to their specific substrates, forming an Enzyme-Substrate Complex (ES). This complex then facilitates the chemical transformation, breaking down to release the enzyme (unchanged) and the new product(s).
The general reaction can be represented as:
Where:
- = Enzyme
- = Substrate
- = Enzyme-Substrate Complex
- = Product
There are two primary models explaining enzyme-substrate interaction:
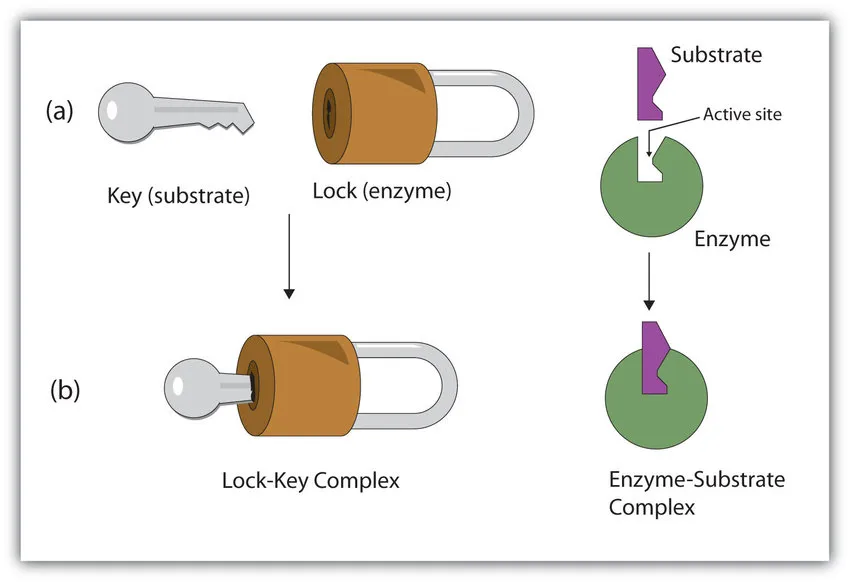
- Lock and Key Model (1894, Emil Fischer):
- This model proposes that enzymes and substrates possess specific, complementary shapes that fit together perfectly, much like a key fits into a specific lock.
- It emphasizes the high specificity of enzymes, where only a certain substrate can bind to a certain enzyme.
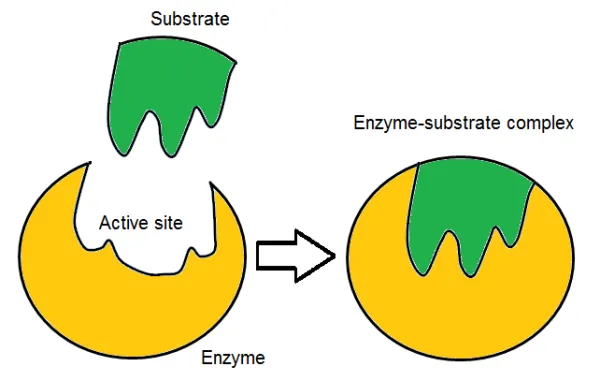
- Induced-Fit Model (1958, Daniel Koshland):
- This model suggests that the active site of an enzyme is not a rigid structure but rather undergoes a conformational change (molds its shape) upon binding with the substrate.
- The substrate "induces" a change in the enzyme's shape to achieve a tighter fit, optimizing the interaction for catalysis. This model is considered more acceptable than the lock and key model as it accounts for the flexibility of enzyme structures.
15.4.2 Factors Affecting Enzyme Activity
Enzymes are highly sensitive to their environment. Any factor that alters the chemistry or three-dimensional shape of an enzyme molecule can significantly impact its function.
i) Temperature
- Optimum Temperature: Enzymes exhibit maximum activity at a specific temperature called the optimum temperature. For example, most animal enzymes function optimally around , while some plant enzymes operate effectively at .
- Effect of Increasing Temperature: Initially, increasing temperature increases the kinetic energy of reactant molecules, leading to more frequent collisions between enzyme and substrate, thus increasing the reaction rate.
- Enzyme Denaturation: However, if the temperature rises too high above the optimum, the intense vibrations of enzyme atoms disrupt the enzyme's delicate globular protein structure. This irreversible process, known as denaturation, leads to a loss of the active site's shape, drastically reducing or completely abolishing enzyme activity.
ii) Substrate Concentration
- Initial Increase in Rate: When there are sufficient enzyme molecules available, increasing the substrate concentration generally increases the rate of reaction. This is because more substrate molecules mean a higher chance of collision with enzyme active sites.
- Saturation (Active Site State): If the enzyme concentration remains constant while substrate concentration continues to increase, a point is reached where all enzyme active sites are continuously occupied (saturated) with substrate molecules. At this point, further increases in substrate concentration will not increase the reaction rate, as there are no free active sites available to bind additional substrate. The enzyme is working at its maximum capacity.
iii) pH
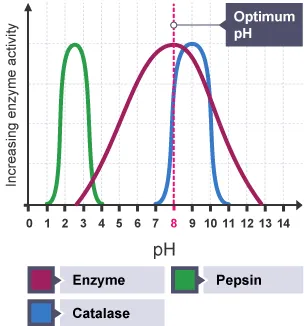
- Optimum pH: Each enzyme has a specific optimum pH range at which it exhibits maximum activity.
- Effect of pH Changes: Significant deviations from the optimum pH (either too acidic or too alkaline) can alter the ionization state of amino acid residues in the enzyme's active site, leading to changes in the enzyme's three-dimensional structure and affecting its ability to bind substrates or catalyze reactions. This can slow down or completely stop enzyme activity.
- Examples:
- Pepsin, an enzyme found in the stomach, is active in highly acidic environments (optimum ).
- Trypsin, an enzyme in the small intestine, is active in alkaline conditions (optimum to ).
15.4.3 Role of Inhibitors in Enzyme-Catalyzed Reactions
Inhibitors are molecules that regulate enzyme-catalyzed reactions by modulating enzyme activity. They are classified into two main types:
-
Competitive Inhibitors:
- Mechanism: These molecules have a structure similar to the enzyme's natural substrate. They compete directly with the substrate for binding to the active site of the enzyme. By binding to the active site, competitive inhibitors prevent the substrate from attaching and forming products.
- Effect: Reduces the rate of reaction by decreasing the effective number of enzyme-substrate interactions. This inhibition can often be overcome by increasing the substrate concentration.
- Examples:
- Sarin (nerve gas): Acts as a competitive inhibitor for acetylcholine (a neurotransmitter). Sarin mimics acetylcholine's structure, binding to its active site and causing acetylcholine to accumulate in synapses, leading to severe physiological effects.
- Malonic acid: Has a structure similar to succinic acid. Succinic acid is normally converted to fumaric acid by the enzyme succinate dehydrogenase. Malonic acid competes with succinic acid for the active site, thereby inhibiting the formation of fumaric acid.
-
Non-Competitive Inhibitors:
- Mechanism: Unlike competitive inhibitors, non-competitive inhibitors do not bind to the active site. Instead, they bind to a different site on the enzyme (an allosteric site), causing a change in the enzyme's overall shape and conformation. This conformational change alters the active site, reducing the enzyme's ability to bind the substrate or catalyze the reaction efficiently.
- Effect: Decreases the maximum reaction rate () of the enzyme, as the enzyme's catalytic efficiency is reduced. This inhibition cannot be overcome by simply increasing the substrate concentration.
- Examples:
- Mercury ions (): Can bind to thiol groups () on enzyme molecules. This binding causes a change in the enzyme's shape and activity, leading to inhibition.
Possible Questions/Answers
-
Q1: What are enzymes? Why are they called biocatalysts? A1: Enzymes are complex protein molecules that act as biological catalysts. They are called biocatalysts because they accelerate (catalyze) the rate of biochemical reactions in living organisms without being consumed in the process, making life-sustaining metabolic activities possible.
-
Q2: How does an enzyme work? A2: Enzymes work by binding specifically to their substrate molecules to form a temporary Enzyme-Substrate (ES) complex. Within this complex, the enzyme facilitates the conversion of the substrate into product(s). After the reaction, the product(s) are released, and the enzyme is regenerated unchanged, ready to catalyze another reaction. This process is often explained by the "Lock and Key" or "Induced Fit" models, which describe how the enzyme's active site interacts with the substrate.
-
Q3: Differentiate between competitive and non-competitive enzyme inhibition. A3:
| Feature | Competitive Inhibitor | Non-Competitive Inhibitor |
|---|---|---|
| Binding Site | Binds to the enzyme's active site. | Binds to a site other than the active site (allosteric site). |
| Similarity | Structurally similar to the substrate. | Structurally dissimilar to the substrate. |
| Mechanism | Competes directly with the substrate for the active site. | Changes the overall shape of the enzyme, altering the active site. |
| Effect | Can be overcome by increasing substrate concentration. | Cannot be overcome by increasing substrate concentration. |
| Example | Sarin (inhibits acetylcholine esterase), Malonic acid (inhibits succinate dehydrogenase). | Mercury ions (), Heavy metal ions. |