10.3 Molecular Carriers or Vectors
This section details the role and characteristics of molecular vectors, particularly plasmids, in recombinant DNA technology. It explains how these vectors are used to carry foreign DNA into host cells for cloning and expression, the essential features that make them effective, and the step-by-step process of creating and selecting for cells containing recombinant DNA.
1. Molecular Vectors
A molecular vector is a DNA molecule used as a vehicle to carry a foreign DNA fragment (gene of interest) into a host cell. The primary purpose is to enable the replication (cloning) and/or expression of the inserted gene within the host.
Examples of Vectors:
- Plasmids: The most common type; small, circular DNA molecules from bacteria.
- Lambda phage DNA: A viral vector.
- Cosmid: A hybrid vector combining features of plasmids and phage DNA.
- Yeast Artificial Chromosomes (YACs) and Bacterial Artificial Chromosomes (BACs): Used for cloning very large DNA fragments.
2. Plasmids as Vectors
Plasmids are small, circular, extrachromosomal DNA molecules found naturally in bacteria and some eukaryotes (e.g., yeast). They replicate independently of the host cell's main chromosome. Naturally, they often carry genes beneficial to the host, such as those for antibiotic resistance and fertility.
Historical Examples:
- pSC-101: Carries a gene for tetracycline resistance.
- pBR322: Carries genes for both tetracycline and ampicillin resistance.
3. Essential Characteristics of a Plasmid Vector
For a plasmid to be an effective vector in genetic engineering, it must possess several key features:
- Origin of Replication (Ori) site: A specific DNA sequence that initiates the self-regulated replication of the plasmid inside the host cell, allowing for the production of multiple copies of the inserted gene.
- Selectable Markers: Genes that confer a trait allowing for the selection of host cells that have successfully taken up the vector. The most common markers are antibiotic resistance genes (e.g., ampicillin or tetracycline resistance).
- Restriction Sites: Specific DNA sequences recognized and cut by restriction endonucleases. A good vector has unique restriction sites (cloning sites) where a foreign gene can be inserted without disrupting essential functions.
- Promoter Regions: DNA sequences that enable the host cell's machinery to transcribe the inserted gene, leading to the production of the desired protein (gene expression).
- Small Size: Smaller plasmids are more easily isolated, manipulated, and inserted into host cells (transformation is more efficient).
- High Copy Number: Some plasmids exist in many copies within a single host cell, leading to a high yield of the cloned DNA or its protein product.
4. Case Study: pBR322 Plasmid
pBR322 was the first widely used, purpose-built cloning vector, created by Bolivar and Rodriguez in 1977.
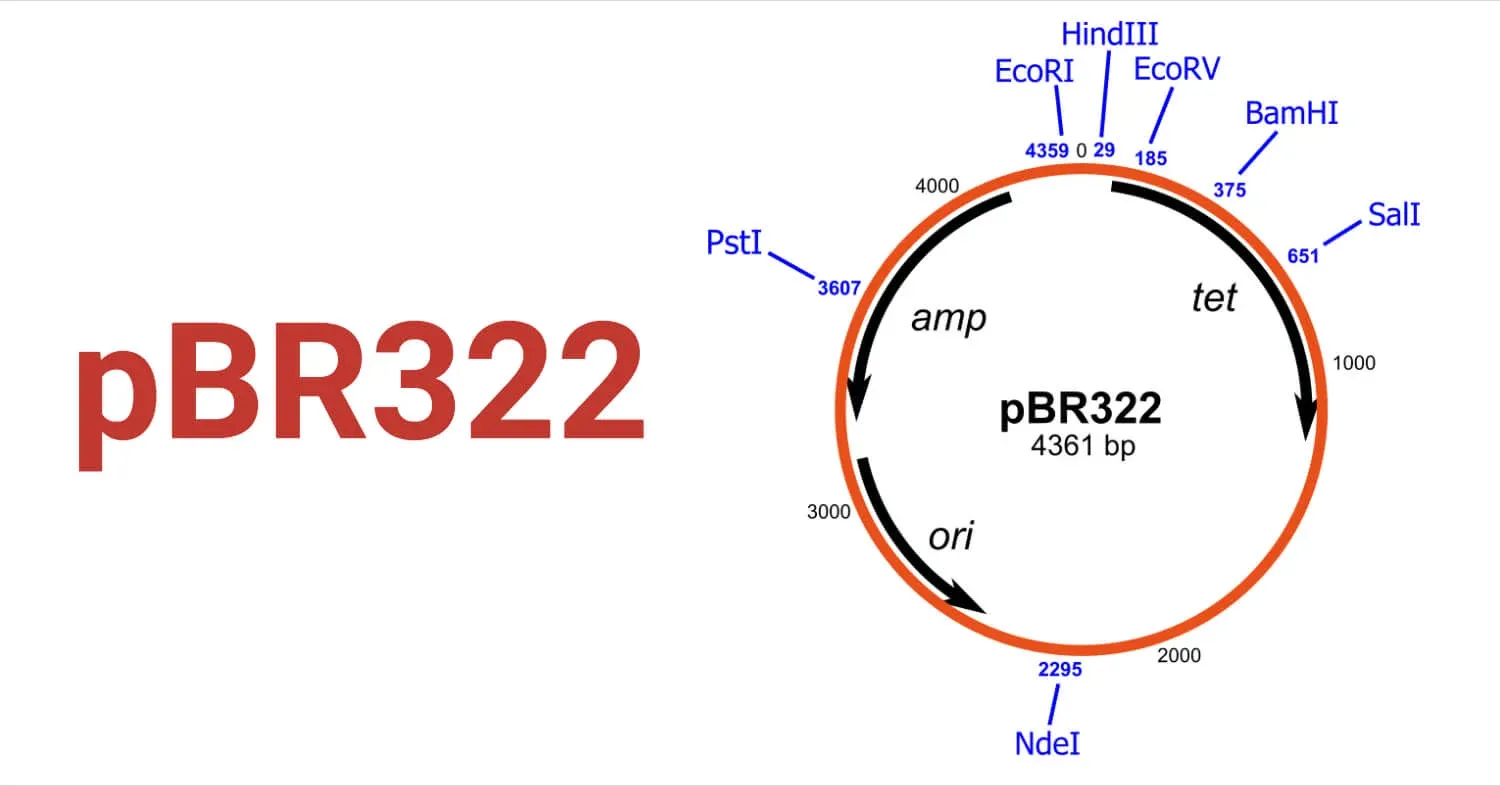
Key Features of pBR322:
- Origin of replication (ori): Allows it to replicate in E. coli.
- Size: Relatively small at 4,361 base pairs (bp), which allows for efficient transformation and insertion of DNA fragments up to ~6 kbp.
- Copy Number: A relatively high copy number of about 15 copies per cell.
- Selectable Markers: Contains two antibiotic resistance genes:
- Ampicillin resistance (Amp<sup>R</sup>)
- Tetracycline resistance (Tet<sup>R</sup>)
- Cloning Sites: Contains several unique restriction sites located within the antibiotic resistance genes. For example, sites for PstI and PvuI are in the ampicillin gene, while sites for BamHI and HindIII are in the tetracycline gene. This allows for a technique called insertional inactivation to screen for recombinant plasmids.
5. Types and Applications of Plasmids
| Types of Plasmid Vectors | Description |
|---|---|
| Cloning Vectors | Designed primarily to clone (make many copies of) a DNA fragment. Contain Ori, markers, and cloning sites. |
| Expression Vectors | Designed for the expression (transcription and translation) of a gene to produce a protein. Include promoters and other regulatory sequences. |
| Shuttle Vectors | Can replicate in multiple host species (e.g., both bacteria and yeast), allowing genes to be moved between different systems. |
| Applications of Plasmids | Description |
|---|---|
| Gene Cloning | Used to make many copies of a gene of interest for study, sequencing, or further manipulation. |
| Protein Production | Expression vectors are used to produce large quantities of proteins like insulin, growth hormone, or enzymes. |
| Gene Therapy | Plasmids can be used as vectors to deliver a correct copy of a faulty gene into the cells of a patient. |
| Vaccine Development | DNA vaccines are often plasmid-based, carrying a gene from a pathogen to stimulate an immune response. |
6. Mechanism of Recombinant Plasmid Formation
The process involves creating a new combination of DNA (recombinant DNA) and introducing it into a host organism.
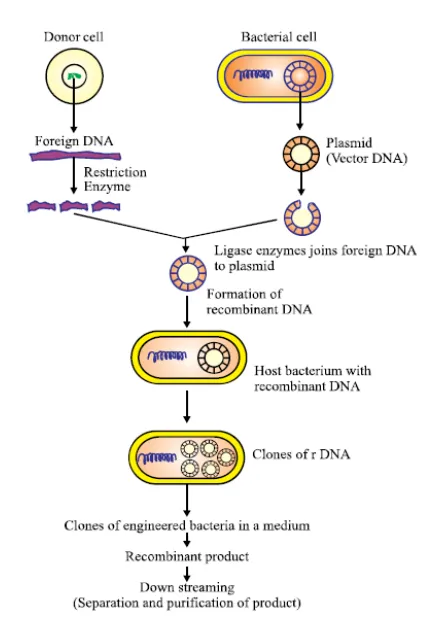
1. Formation of Recombinant Plasmid
- Isolation: The gene of interest and the plasmid vector are isolated and purified.
- Digestion: Both the gene and the plasmid are cut with the same restriction endonuclease. This creates compatible, complementary sticky ends on both DNA fragments.
- Ligation: The gene of interest and the cut plasmid are mixed together. The enzyme DNA ligase acts as a molecular glue, forming phosphodiester bonds to permanently join the gene into the plasmid. The resulting hybrid DNA molecule is called a recombinant plasmid.
2. Transformation of Expression System
- Expression System: A host cell or organism (e.g., E. coli bacteria) is chosen to receive the recombinant DNA. Ideal hosts have a short generation time and are easy to grow.
- Transformation: The process of introducing the recombinant plasmid into the host cell. This is often done by treating the bacterial cells with calcium chloride (CaCl₂), which makes their membranes more permeable to DNA.
3. Identification of Transformed Clones (Selection)
After transformation, the mixture contains three types of cells: untransformed cells, cells with the original (non-recombinant) plasmid, and cells with the recombinant plasmid.
- Selection: The cells are grown on a medium containing an antibiotic corresponding to the plasmid's resistance gene.
- Outcome:
- Untransformed cells lack the plasmid and its resistance gene, so they are killed by the antibiotic.
- Transformed cells (containing the plasmid) survive and multiply, forming a clone. These clones can then be used to produce the protein of interest or to isolate large quantities of the cloned gene.