10.2 Molecular Scissors: Restriction Endonuclease
Restriction Endonucleases: These are enzymes that recognize specific DNA sequences and cleave the phosphodiester bonds of both DNA strands within that sequence. They are crucial tools for cutting a gene of interest during genetic recombination.
Natural Function: A Bacterial Defense System
- Restriction endonucleases serve as a natural defense mechanism in bacteria against invading viruses (bacteriophages).
- They "restrict" viral growth by digesting (cutting up) the viral DNA at specific recognition sites.
- The bacterial cell's own DNA is protected from its own restriction enzymes through a process called methylation.
Host DNA Protection: Methylation
- Bacteria add a methyl group () to nucleotides within their own restriction sites.
- This modification is carried out by an enzyme called methyltransferase.
- The added methyl groups block the restriction endonuclease from binding and cleaving the host's DNA, while the un-methylated viral DNA remains vulnerable.
Nomenclature of Restriction Enzymes
-
Enzymes are named after the genus, species, and strain of the bacteria from which they were isolated.
-
Examples:
- EcoRI: Isolated from Escherichia coli (strain RY13, I indicating the first enzyme identified).
- Hind-II & Hind-III: Isolated from Haemophilus influenzae (strain d).
- XhoI: Isolated from Xanthomonas holcicola.
Recognition Sites and Types of Cuts
- Palindromic Sequences: Restriction enzymes recognize specific sequences of 4 to 8 base pairs that are palindromic. This means the nucleotide sequence on one strand reads the same in the opposite direction on the complementary strand (e.g., 5'-GAATTC-3' on one strand and 3'-CTTAAG-5' on the other).
-
Types of Cuts:
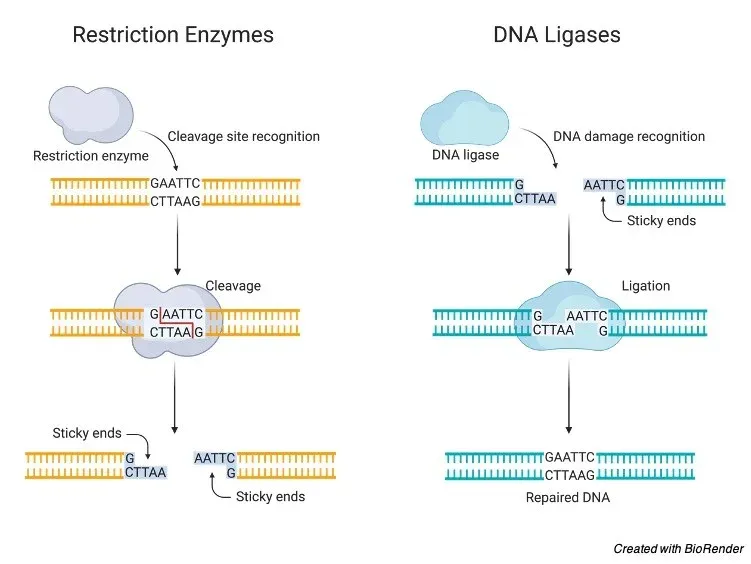
- Staggered Cuts (Sticky Ends): The enzyme cuts the two DNA strands at different points within the recognition site, creating single-stranded overhangs called sticky ends. These ends are "sticky" because they can easily form hydrogen bonds with complementary sequences.
- Example (EcoRI): Produces sticky ends at the 5' end of each strand.
- Staggered Cuts (Sticky Ends): The enzyme cuts the two DNA strands at different points within the recognition site, creating single-stranded overhangs called sticky ends. These ends are "sticky" because they can easily form hydrogen bonds with complementary sequences.
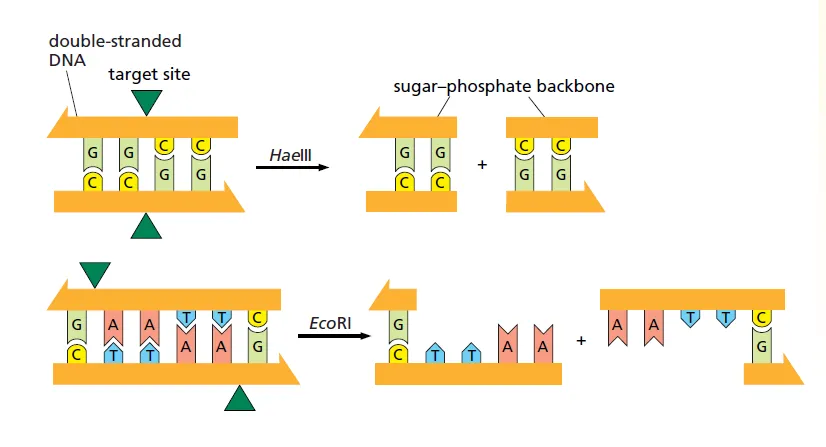
- Blunt Cuts: The enzyme cuts both DNA strands at the same point, resulting in fragments with no overhangs. These are called blunt ends.
- Example (HaeIII): Cuts directly in the middle of its recognition sequence (GGCC).
Possible Questions/Answers
Q: Why don't restriction endonucleases destroy the DNA of their host bacterium? A: The host bacterium protects its own DNA by adding methyl groups () to the nucleotides in its recognition sequences. This process, called methylation, blocks the restriction enzyme from binding and cutting the host's DNA.
Q: Why are restriction enzymes that create sticky ends more useful in genetic engineering? A: Sticky ends are single-stranded overhangs that are complementary to each other. This allows a DNA fragment (like a gene of interest) and a cloning vector (like a plasmid) cut with the same enzyme to easily join together (anneal) through base pairing, which greatly facilitates the process of creating recombinant DNA.